Available with Network Analyst license.
Summary
Constructs a spatial weights matrix file (.swm) using a Network dataset, defining feature spatial relationships in terms of the underlying network structure.
Illustration
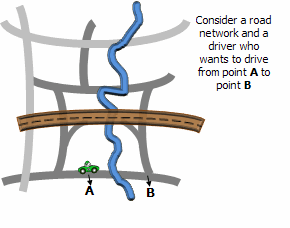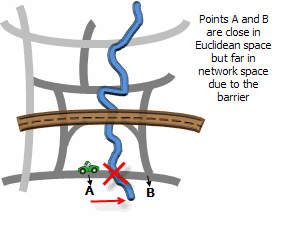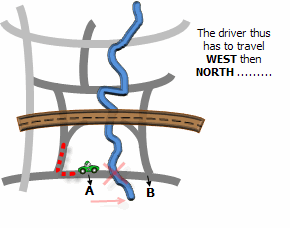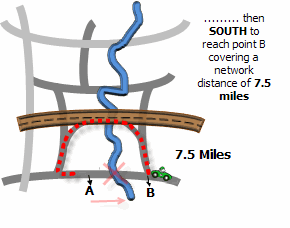
Usage
Output from this tool is a spatial weights matrix file (.swm). Tools that require you to specify a Conceptualization of Spatial Relationships option will accept a spatial weights matrix file; select GET_SPATIAL_WEIGHTS_FROM_FILE for the Conceptualization of Spatial Relationships parameter and, for the Weights Matrix File parameter, specify the full path to the spatial weights file created using this tool.
This tool was designed to work with point Input Feature Class data only.
A spatial weights matrix quantifies the spatial relationships that exist among the features in your dataset. Many tools in the Spatial Statistics toolbox evaluate each feature within the context of its neighboring features. The spatial weights matrix file defines those neighbor relationships. For this tool, neighbor relationships are based on the time or distance between features, in the case where travel is restricted to a network. For more information about spatial weights and spatial weights matrix files, see Spatial_weights.
-
The Unique ID field is linked to feature relationships derived from running this tool. Consequently, the Unique ID values must be unique for every feature and typically should be in a permanent field that remains with the feature class. If you don't have a unique ID field, you can create one by adding a new integer field (Add Field) to your feature class table and calculating the field values to be equal to the FID or OBJECTID field (Calculate Field). Because the FID and OBJECTID field values may change when you copy or edit a feature class, you cannot use these fields directly for the Unique ID parameter.
The Maximum Number of Neighbors parameter for this tool specifies the exact number of neighbors that will be associated with each feature. The Impedance Cutoff overrides the number of neighbors parameter, so some features may have fewer neighbors if the number of neighbors specified cannot be found within the cutoff distance/time.
You can define spatial relationships using the hierarchy in the network dataset, if it has one, by checking the Use Hierarchy in Analysis parameter. The hierarchy classifies network edges into primary, secondary, and local roads. When using the hierarchy of the network to create spatial relationships among features, preference will be given to travel on primary roads more than secondary roads and secondary roads more than local roads.
This tool does not honor the output coordinate system environment setting. All feature geometry is projected to match the spatial reference associated with the Network Dataset prior to analysis. The resultant spatial weights matrix file created by this tool will reflect spatial relationships defined using the Network Dataset spatial reference. It is recommended that when performing analyses using a network spatial weights matrix file, the input feature class be projected to match the coordinate system of the network dataset used to create the network SWM.
Syntax
GenerateNetworkSpatialWeights_stats (Input_Feature_Class, Unique_ID_Field, Output_Spatial_Weights_Matrix_File, Input_Network, Impedance_Attribute, {Impedance_Cutoff}, {Maximum_Number_of_Neighbors}, {Barriers}, {U-turn_Policy}, {Restrictions}, {Use_Hierarchy_in_Analysis}, {Search_Tolerance}, {Conceptualization_of_Spatial_Relationships}, {Exponent}, {Row_Standardization})| Parameter | Explanation | Data Type |
Input_Feature_Class | The point feature class for which network spatial relationships among features will be assessed. | Feature Class |
Unique_ID_Field | An integer field containing a different value for every feature in the input feature class. If you don't have a Unique ID field, you can create one by adding an integer field to your feature class table and calculating the field values to equal the FID or OBJECTID field. | Field |
Output_Spatial_Weights_Matrix_File | The output network spatial weights matrix (.swm) file. | File |
Input_Network | The network dataset for which spatial relationships among features in the input feature class will be defined. | Network Dataset Layer |
Impedance_Attribute | The type of cost units to use as impedance in the analysis. | String |
Impedance_Cutoff (Optional) |
Specifies a cutoff value for INVERSE and FIXED conceptualizations of spatial relationships. Enter this value using the units specified by the Impedance_Attribute parameter. A value of zero indicates that no threshold is applied. When this parameter is left blank, a default threshold value is computed based on input feature class extent and the number of features. | Double |
Maximum_Number_of_Neighbors (Optional) | An integer reflecting the maximum number of neighbors to find for each feature. | Long |
Barriers (Optional) | The name of a point feature class with features representing blocked intersections, road closures, accident sites, or other locations where travel is blocked along the network. | Feature Layer |
U-turn_Policy (Optional) | Specifies optional U-turn restrictions.
| String |
Restrictions [Restriction,...] (Optional) | A list of restrictions. Check the restrictions to be honored in spatial relationship computations. | String |
Use_Hierarchy_in_Analysis (Optional) | Specifies whether or not to use a hierarchy in the analysis.
| Boolean |
Search_Tolerance (Optional) | The search threshold used to locate features in the Input_Feature_Class onto the network dataset. This parameter includes a search value and the units for the tolerance. | Linear unit |
Conceptualization_of_Spatial_Relationships (Optional) | Specifies how the weighting associated with each spatial relationship is specified.
| String |
Exponent (Optional) |
Parameter for the INVERSE Conceptualization_of_Spatial_Relationships calculation. Typical values are 1 or 2. Weights drop off quicker with distance as this exponent value increases. | Double |
Row_Standardization (Optional) | Row standardization is recommended whenever feature distribution is potentially biased due to sampling design or to an imposed aggregation scheme.
| Boolean |
Code sample
GenerateNetworkSpatialWeights example 1 (Python window)
The following Python window script demonstrates how to use the GenerateNetworkSpatialWeights tool.
import arcpy
arcpy.env.workspace = "c:/data"
arpcy.GenerateNetworkSpatialWeights_stats("Hospital.shp", "MyID","network6Neighs.swm",
"Streets_ND","MINUTES", 10, 6, "#",
"ALLOW_UTURNS","#", "USE_HIERARCHY",
"#", "INVERSE", 1,"ROW_STANDARDIZATION")
GenerateNetworkSpatialWeights example 2 (stand-alone Python script)
The following stand-alone Python script demonstrates how to use the GenerateNetworkSpatialWeights tool.
# Create a Spatial Weights Matrix based on Network Data
# Import system modules
import arcpy
# Set the geoprocessor object property to overwrite existing output
arcpy.gp.overwriteOutput = True
# Check out the ArcGIS Network Analyst extension (required for the Generate Network Spatial Weights tool)
arcpy.CheckOutExtension("Network")
# Local variables...
workspace = r"C:\Data"
try:
# Set the current workspace (to avoid having to specify the full path to the feature classes each time)
arcpy.env.workspace = workspace
# Create Spatial Weights Matrix based on Network Data
# Process: Generate Network Spatial Weights...
nwm = arcpy.GenerateNetworkSpatialWeights_stats("Hospital.shp", "MyID",
"network6Neighs.swm", "Streets_ND",
"MINUTES", 10, 6, "#", "ALLOW_UTURNS",
"#", "USE_HIERARCHY", "#", "INVERSE",
1, "ROW_STANDARDIZATION")
# Create Spatial Weights Matrix based on Euclidean Distance
# Process: Generate Spatial Weights Matrix...
swm = arcpy.GenerateSpatialWeightsMatrix_stats("Hospital.shp", "MYID",
"euclidean6Neighs.swm",
"K_NEAREST_NEIGHBORS",
"#", "#", "#", 6)
# Calculate Moran's Index of Spatial Autocorrelation for
# average hospital visit times using Network Spatial Weights
# Process: Spatial Autocorrelation (Morans I)...
moransINet = arcpy.SpatialAutocorrelation_stats("Hospital.shp", "VisitTime",
"NO_REPORT", "GET_SPATIAL_WEIGHTS_FROM_FILE",
"EUCLIDEAN_DISTANCE", "NONE", "#",
"network6Neighs.swm")
# Calculate Moran's Index of Spatial Autocorrelation for
# average hospital visit times using Euclidean Spatial Weights
# Process: Spatial Autocorrelation (Morans I)...
moransIEuc = arcpy.SpatialAutocorrelation_stats("Hospital.shp", "VisitTime",
"NO_REPORT", "GET_SPATIAL_WEIGHTS_FROM_FILE",
"EUCLIDEAN_DISTANCE", "NONE", "#",
"euclidean6Neighs.swm")
except:
# If an error occurred when running the tool, print out the error message.
print arcpy.GetMessages()
Environments
Licensing information
- ArcGIS Desktop Basic: Requires Network Analyst
- ArcGIS Desktop Standard: Requires Network Analyst
- ArcGIS Desktop Advanced: Requires Network Analyst
Related topics
- An overview of the Modeling Spatial Relationships toolset
- Spatial Autocorrelation (Global Moran's I)
- High/Low Clustering (Getis-Ord General G)
- Cluster and Outlier Analysis (Anselin Local Moran's I)
- Hot Spot Analysis (Getis-Ord Gi*)
- Grouping Analysis
- Spatial weights
- Modeling spatial relationships
- Generate Spatial Weights Matrix
- What is a network dataset?